A Selection of eDNA Publications
Please note this collection is not exhaustive, and there are many eDNA publications world-wide. If you would like to add your own paper or have other publication suggestions, please fill out the Publication Submission form at the bottom of the page.

eDNA Monitoring for Deep-Sea Sharks: Minimum Standards for the Reopened Maldives Gulper Shark Fishery
- (Yin Cheong Aden Ip,
- Aquatic Conservation: Marine and Freshwater Ecosystems)
The Maldives reopened a collapsed gulper shark fishery in November 2025 without stock assessment or recovery evidence. Applying the precautionary approach requires three minimum standards for managing reopened or proposed fisheries: transparent recovery assessment, independent population monitoring and funded operational plans with decision rules. I propose that environmental DNA (eDNA) offers a non-lethal, scalable monitoring tool to support evidence-based fishery decisions for threatened deep-sea sharks.

Understanding practical barriers to the global adoption of environmental DNA (eDNA) methods, tools, and standards
- (Shana Hirsch, Yin Cheong Aden Ip, Pedro F. P. Brandão-Dias, Elizabeth Andruszkiewicz Allan & Ryan Kelly ,
- BMC Research Notes)
Environmental DNA (eDNA) is a rapidly emerging data source with the potential to support environmental monitoring and biodiversity conservation around the world. Current efforts to standardize eDNA methods and reporting are aimed at strengthening credibility and supporting adoption. In doing this, however, researchers must be mindful of diverse capacities and ecological contexts both regionally and around the world. The objective of our research is to understand how international standards for eDNA may support or hinder the uptake of eDNA methods and tools for conservation and biodiversity work. This was accomplished through two interactive workshops that brought together eDNA researchers and practitioners from around the world to surface broad and specific barriers to uptake of eDNA methods and tools.
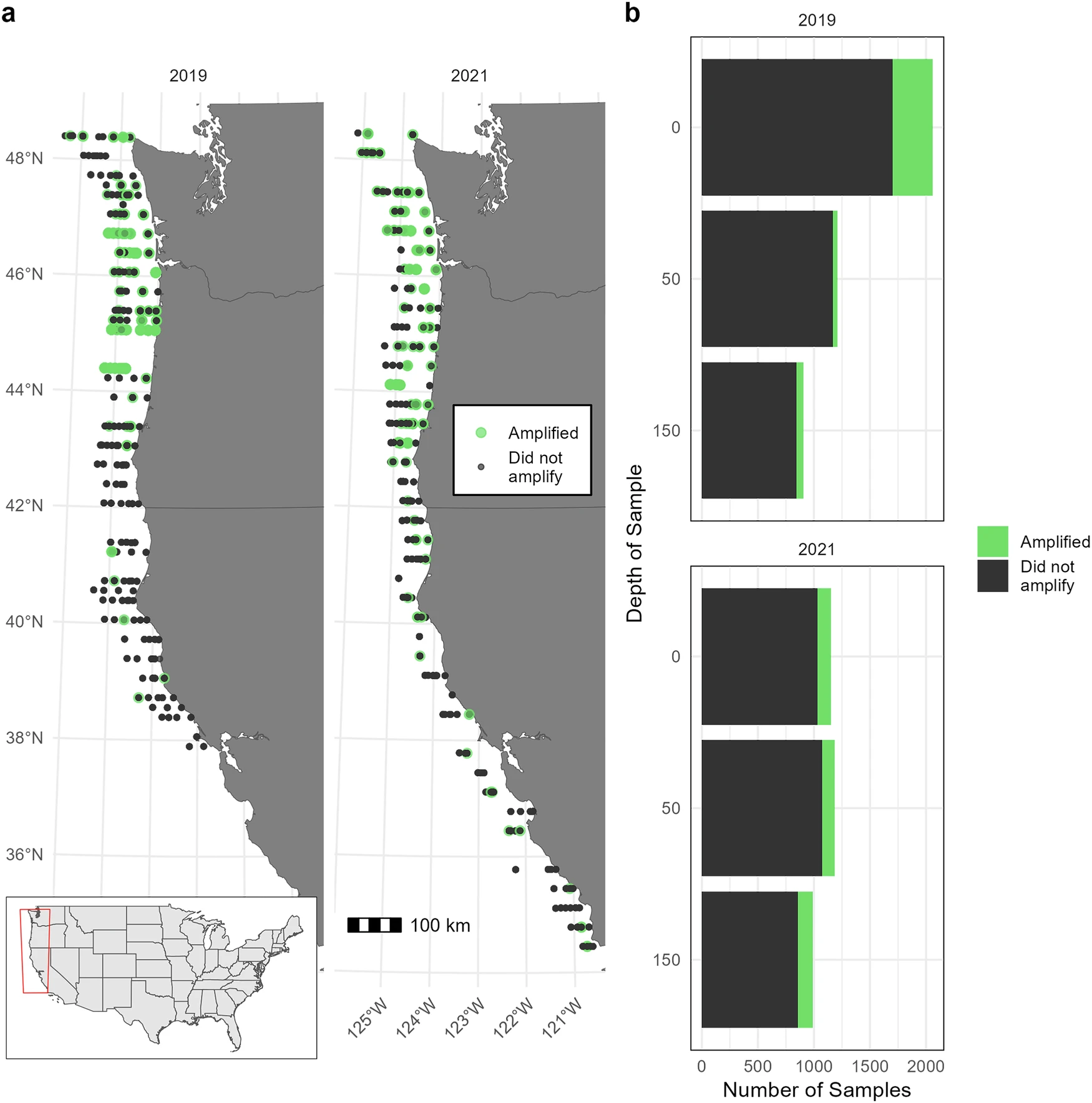
Mapping the marine distribution of eulachon (Thaleichthys pacificus) in the Northeast Pacific using environmental DNA
- (Owen R. Liu, Andrew O. Shelton, Ana Ramón-Laca, Krista M. Nichols, Eric J. Ward, Elizabeth M. Phillips, Jeannette E. Zamon, Abigail Wells & Ryan P. Kelly ,
- Communications Biology)
Rare species are difficult to observe in the wild, particularly in the ocean where large spatial scales and accessibility hinder effective sampling. Environmental DNA (eDNA) is a non-destructive, scalable sampling method with the potential to inform the distribution of rare species in marine ecosystems. We sample eDNA within the California Current ecosystem to estimate the distribution of eulachon (Thaleichthys pacificus), a threatened anadromous smelt ranging along the coastal Northeast Pacific. We amplify eulachon DNA from thousands of water samples collected at night across two years and more than 200,000 square kilometers along the U.S. west coast. We then use spatiotemporal models to derive quantitative estimates of eulachon DNA across space, depth, and time relative to environmental covariates. We find that eulachon DNA has a distribution weighted towards the ocean surface, spatially associated with major river mouths and productive offshore banks. Temperature and prey density are key covariates, with eulachon more likely to be found in warmer waters with higher prey concentrations. We discuss how our results can augment the information currently used in eulachon recovery planning, and describe the wide applicability of our statistical models for estimating distribution and abundance for other species of conservation concern.

Estimating Organism Abundance Using Within-Sample Haplotype Frequencies of eDNA Data
- (Pedro F. P. Brandão-Dias, Gledis Guri, Megan R. Shaffer, Elizabeth Andruszkiewicz Allan, Ryan P. Kelly,
- Molecular Ecology Resources)
Environmental DNA (eDNA) provides powerful insights into species presence and community composition but remains limited in its capacity to infer species abundance or population structure. Here, we show that the deviation between within-sample haplotype frequencies and the overall population-level haplotype frequencies can be used to estimate the number of individual contributors to a given sample. We first establish the theoretical framework for approximating population haplotype frequencies directly from eDNA data, enabling application even in the absence of tissue-derived references. Building on this foundation, we introduce a maximum likelihood estimator to infer the number of contributors and assess its performance through simulations spanning a range of haplotype frequency distributions and noise scenarios. These approaches assume that all samples are drawn from a single, panmictic population. We find that accurate estimates are attainable when haplotypes are sufficiently variable, population frequencies are well-characterised, and samples are large enough to capture frequency deviations. By bridging population genetic theory and eDNA, our method complements existing molecular approaches and offers a novel path towards quantifying abundance from eDNA metabarcoding data.
Submit Your Publication
The eDNA Collaborative welcomes publication suggestions, yours or someone else’s, for addition to this page. Please submit your suggestion using the form provided. Be sure to include your reason for the recommendation in the “Justification” field.